Limba
|
D5456, Pico Methyl-Seq™ Library Prep Kit (25 preps.)
10.079,30 RON
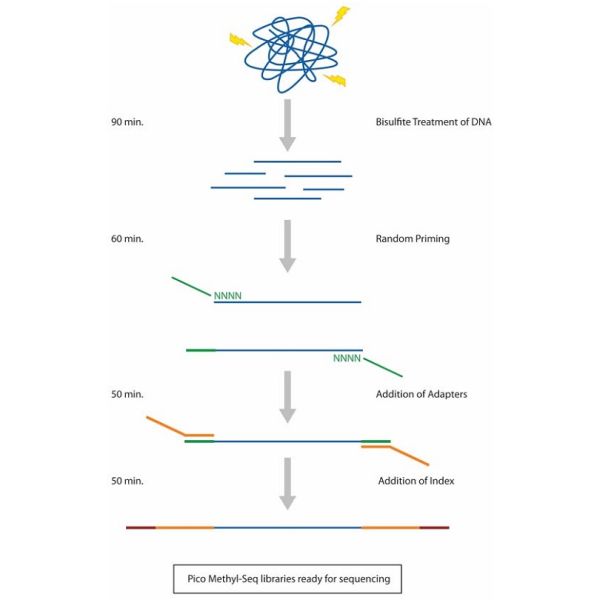
The Pico Methyl-Seq Library Prep Kit provides a streamlined workflow for making WGBS libraries. Input DNA is randomly fragmented during the initial bisulfite treatment step followed by three rounds of amplification with uniquely designed primers. The procedure can accommodate as little as 10 pg input DNA (including that derived from FFPE samples), making it ideal for methylation analysis of precious, limited, and target-enriched samples.
SKU
ZR_D5456
HIGHLIGHTS
- All-inclusive kit for bisulfite conversion followed by Whole Genome Bisulfite Sequencing (WGBS) library preparation.
- Accommodates ultra-low DNA input and compatible with FFPE samples.
- Simple, ligation- and gel-free workflow can be completed in a few hours.
DESCRIPTION
TECHNICAL SPECIFICATIONS
| Equipment | Microcentrifuge, thermocycler |
|---|---|
| Input | 10 pg - 100 ng |
| Sample Source | The protocol is designed for 10 pg – 100 ng genomic DNA input. DNA should be free of enzymatic inhibitors and can be suspended in water, TE, or a low salt buffer. DNA with low 260/280 or 260/230 ratios should be purified prior to processing using the Genomic DNA Clean & Concentrator (D4010). |
| Sequencing Compatibility | This kit produces libraries with Illumina TruSeq Adapters that can be sequenced on any of the Illumina sequencing platforms. |
| Supplemental Info |
| Price | 8.470,00 RON (preturile sunt fara TVA) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Description |
HIGHLIGHTS
DESCRIPTIONTECHNICAL SPECIFICATIONS
|
||||||||||
| Manufacturer | Zymo Research |

 English
English

![D5455, Pico Methyl-Seq™ Library Prep Kit (10 preps.) -[Includes Room Temp Item & Dry Ice Item]](https://biozyme.ro/media/catalog/product/cache/473f3ad9dd2347c84e0ead05c074d48c/d/5/d5455_66_2_nv2mqtm4sj7ovc8p.jpg)
